mainmenu
Center for Virome and Applied Platform
Experimental systems for modeling virus& host interactions
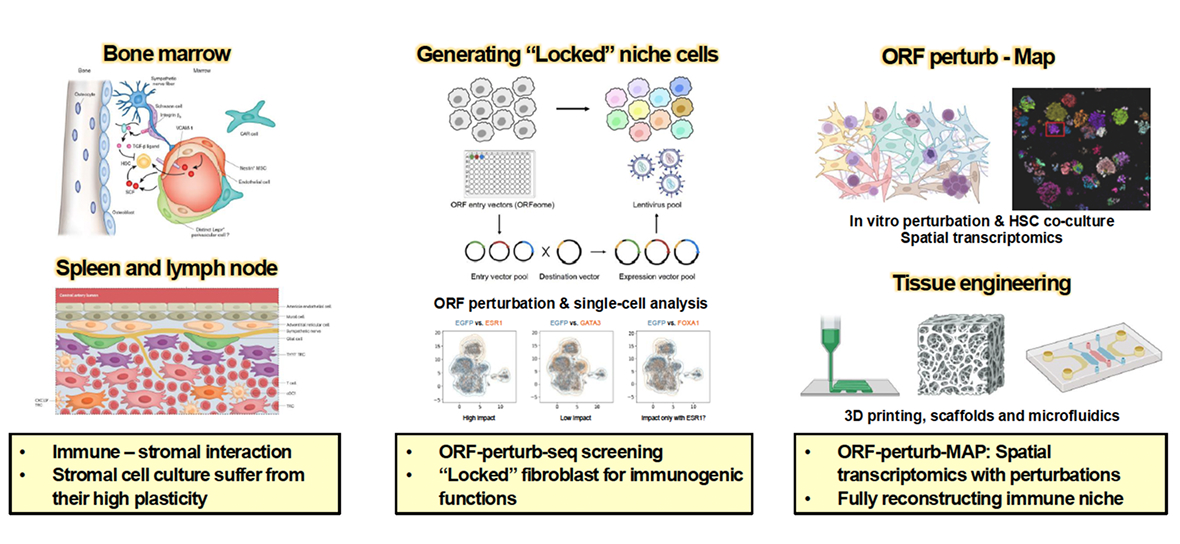
Computational models of virus–host interactions require experimental systems that can reproduce key features of human immune responses. To address this challenge, we develop experimental platforms that integrate epithelial, stromal, and immune components within engineered tissue environments.
Our approach combines organoid-based models with stromal and immune cell engineering to reconstruct physiologically relevant tissue niches. Using perturbation-based single-cell profiling, we map gene regulatory networks that govern stromal–immune interactions and use these insights to engineer stable cellular environments for infection studies. These systems enable controlled investigation of viral tropism, immune activation, and tissue responses, providing an experimental framework to test and refine computational predictions.