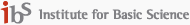mainmenu
Center for Virome and Applied Platform
Modeling host tissue susceptibility to viral infection
Viral infection begins with interactions between viral proteins and host cellular environments that vary across tissues and cell types. To understand these processes, we construct large-scale host tissue atlases using single-cell and spatial transcriptomics data across diverse organs and physiological conditions.
By integrating host receptor expression, cellular states, and tissue microenvironmental features, we develop computational models that predict viral entry pathways and tissue susceptibility. These datasets also serve as the basis for developing machine learning models that capture general patterns of virus–host interactions across tissues. This work builds upon our previous efforts in large-scale single-cell integration and atlas construction to provide a systematic framework for understanding viral tropism.