mainmenu
Center for Virome and Applied Platform
Overview
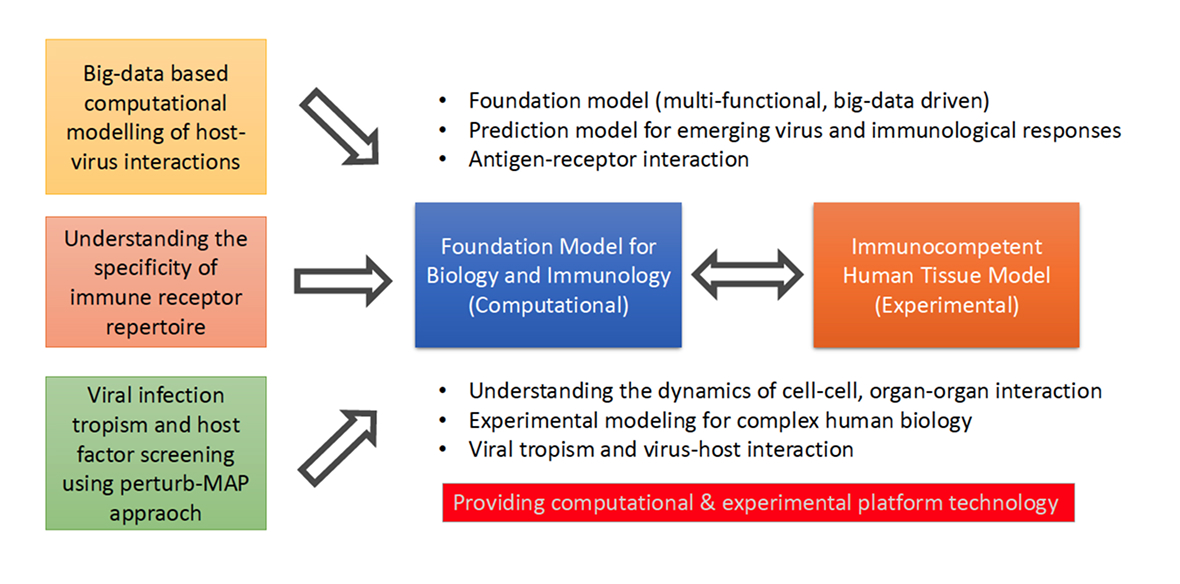
Viral infections emerge from complex interactions between viruses and the host immune system across multiple biological scales, ranging from molecular recognition to tissue-level immune responses. Understanding these processes requires integrating diverse layers of biological information, including cellular states, immune receptor repertoires, and spatial organization of tissues. Our research aims to build a systematic framework for studying virus–host interactions by combining large-scale single-cell and spatial omics datasets with advanced computational modeling.
A central objective of our work is to develop an integrative, data-driven representation of host immunity that captures the diversity of human tissues and immune responses. Building upon our previous studies in large-scale single-cell atlas construction and immune diversity analysis, we integrate multi-organ transcriptomic datasets, immune receptor repertoires, and spatially resolved infection data to uncover the principles governing viral tropism and immune recognition.
In parallel, we aim to develop computational models that can generalize across biological systems. By leveraging recent advances in machine learning and foundation models for biological data, we seek to construct predictive frameworks that connect host cellular states, immune receptor specificity, and viral protein interactions. These models are complemented by experimental systems that allow systematic validation of predicted virus–host interactions. Through the integration of large-scale data analysis and experimental modeling, our research aims to establish a predictive and mechanistic understanding of viral infection and host immunity, providing a foundation for future strategies in antiviral therapeutics and vaccine development.